In other words, due to more accumulated mutations, it appears as if less information is carried by the DNA sequences of AT rich microbes compared to GC rich microbes. An organism's DNA sequence that has been subjected to numerous random mutations is assumed to possess less information than the DNA of an organism under strong selective pressure. the signal-to-noise ratio is lower in AT rich genomes. GC content has also been associated with genome wide rates of mutation, where organisms of low GC content tend to have more random genomes than GC rich ones, i.e. the more GC rich an organism is the more compositionally similar it tends to be with its plasmid(s). In fact, genomic signatures based methods reveal prokaryotic plasmid-host similarity to correlate with genomic GC content, i.e. Plasmids on the other hand, although sharing some similarities with their hosts, have a more different DNA composition than what would be expected compared to the hosts chromosome. Based on findings from genomic signatures (and analyses of CRISPSs in bacteria ), phages, and viruses in general, have been found to co-evolve with their hosts. Mathematical models predict plasmids to be the predominant means of genetic variation among bacteria. 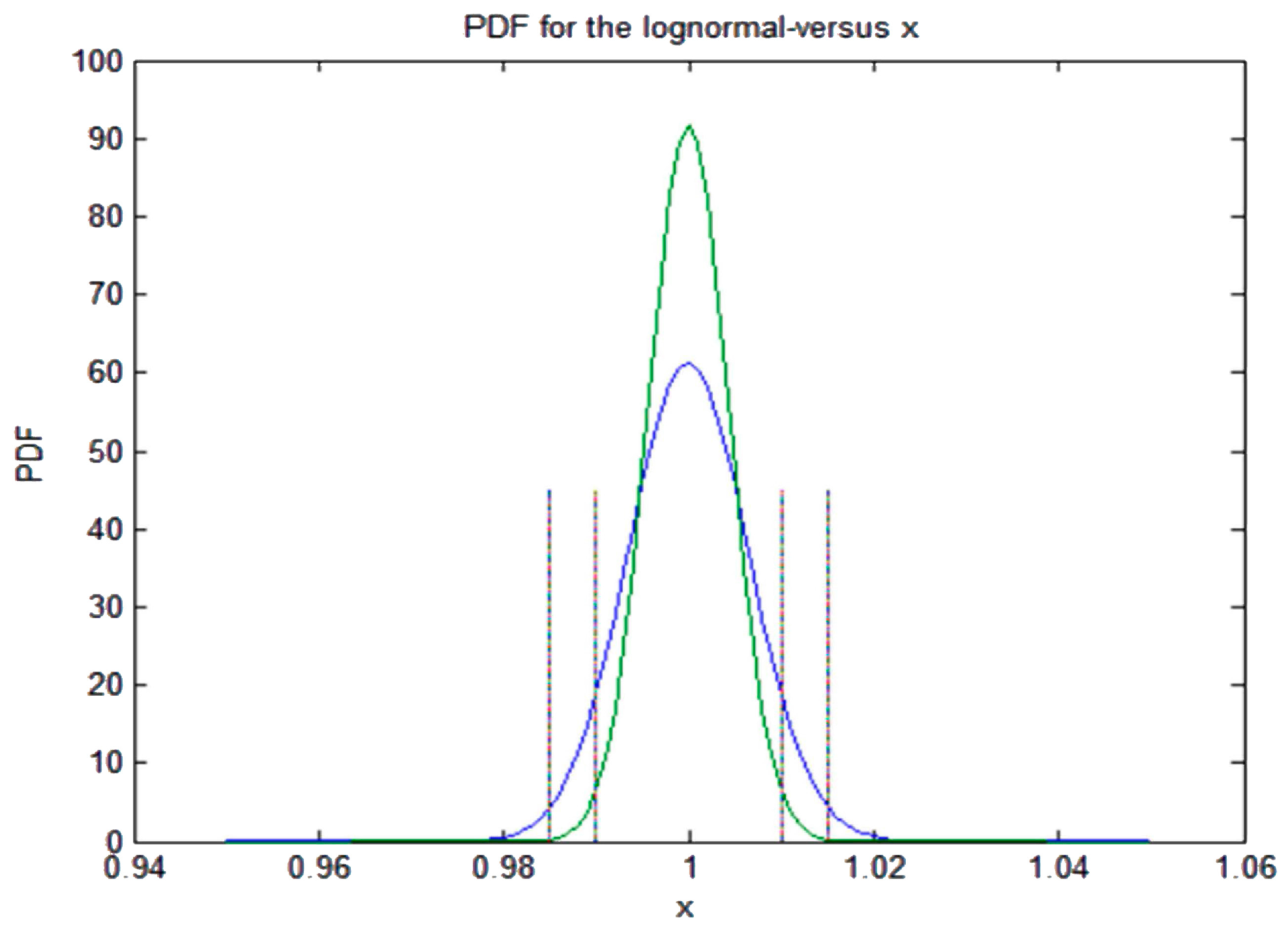
However, this exchange is not entirely restrictive with low frequency transfer occurring between chromosomes on one hand and plasmids and phages on the other. chromosomes exchange DNA with chromosomes, plasmids with plasmids, and phages with phages. Genetic material is most readily exchanged between related genetic elements, i.e. GIs originate from different sources including plasmids and phages (prophages) and carry traits that have important biological phenotypes such as virulence determinants and antibiotic resistance genes. Such putatively horizontally transferred regions are termed Genomic Islands (GI). In addition to plasmids and phages, regions within the bacterial chromosomes are assumed to have been horizontally acquired. Horizontal gene transfer in microbial communities has been recognized as a key driver of evolutionary change in microbes. The rate at which amelioration of horizontally acquired DNA occurs within the chromosome is likely to account for the small differences between chromosomes and stably incorporated GIs compared to the transient or independent replicons such as phages and plasmids. 
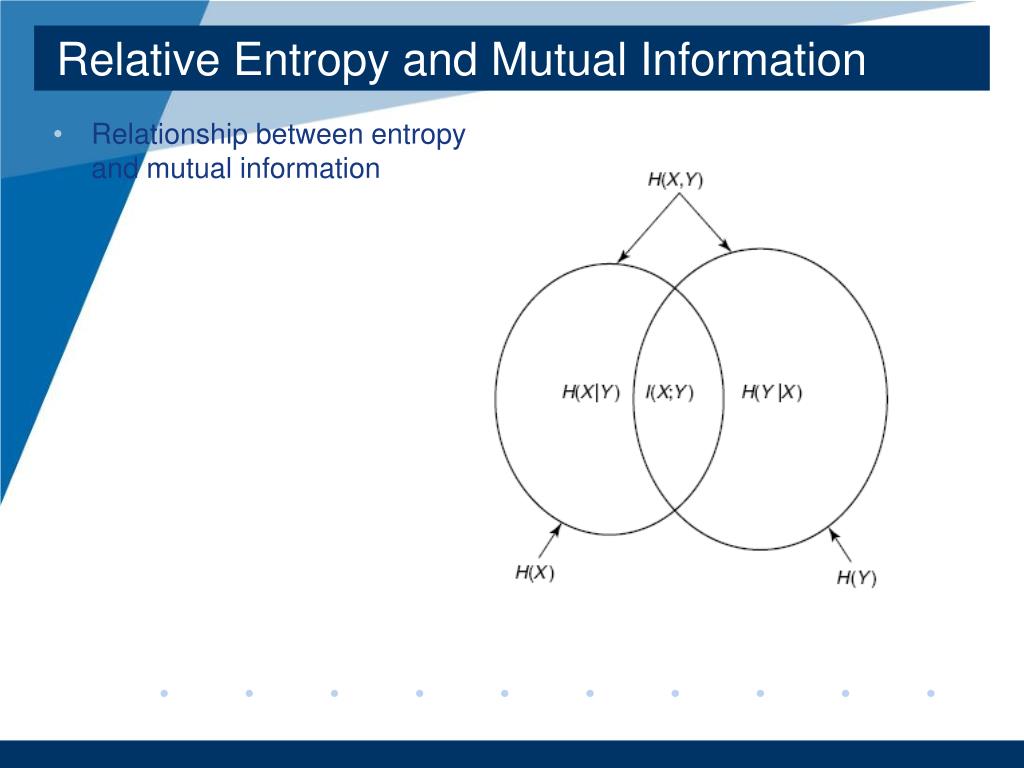
We argue that relative entropy differences reflect how plasmids, phages and GIs interact with microbial host chromosomes and that all these biological entities are, or have been, subjected to different selective pressures. tuberculosis when measured on a shared set of highly conserved genes. Relative entropy was also found to be lower in the obligate intracellular Mycobacterium leprae than in the related M. There was an association between relative entropy and AT content in chromosomes, phages, plasmids and GIs with the strongest association being in phages. Relative entropy was highest in bacterial chromosomes and had the sequence chromosomes > GI > phage > plasmid. Relative entropy was estimated using the Kullback-Leibler measure. We analyzed the differences in information capacity between prokaryotic chromosomes, genomic islands (GI), phages, and plasmids. We sought to assess whether the concept of relative entropy (information capacity), could aid our understanding of the process of horizontal gene transfer in microbes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
